Nanotechnology innovations record molecular geometry and advance molecular robotics
In a pair of recent research papers, work by Thomas Schaus contributes to the development of new nanotechnology tools. Schaus was a JCC Fellow from 2011-2014 in Peng Yin’s lab at Harvard University’s Wyss Institute of Biologically Inspired Engineering.
“These projects began when Peng challenged me to record the organization of macromolecular structures, in as complete a manner as possible, with pieces of DNA,” says Schaus. “There have long been several mechanisms for recording protein-protein interactions and similar structures, but they tend to be ensemble techniques – either inadequately sensitive, or simply unable make more than one measurement from each molecular complex of interest.”
 Optical microscopes and Cryo-EM are two of the techniques that have been used to see static snapshots of molecular structures. These approaches are often successful but have limitations in labeling, resolution, and imaging conditions. Biochemical approaches also aim to reveal molecular structures in detail, but typically destroy the structures in the process, thereby missing important information that would facilitate whole-molecule reconstruction.
Optical microscopes and Cryo-EM are two of the techniques that have been used to see static snapshots of molecular structures. These approaches are often successful but have limitations in labeling, resolution, and imaging conditions. Biochemical approaches also aim to reveal molecular structures in detail, but typically destroy the structures in the process, thereby missing important information that would facilitate whole-molecule reconstruction.
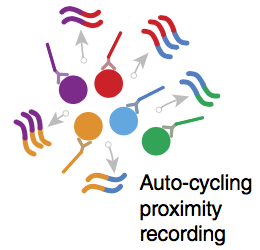
“Generating much more data from a given sample would improve measurement reliability and precision in the short term, and is essential for the longer-term goal of deciphering structures and networks with single-molecule resolution,” Schaus explains. “With these goals in mind we developed APR (Auto-cycling Proximity Recording), in which targets are labeled with unique DNA barcodes, and barcode identities are repeatedly copied in pairs to make DNA ‘records’ of target proximity (see animation). Because labels are copied instead of used up in an irreversible reaction (e.g., ligation), the cycle can proceed autonomously, at physiological conditions, to generate multiple records from any pair of combinations in the same sample. In this paper, we demonstrated cycling recording and ensemble reconstructions of complex geometrical arrangements, as well as repeat measurements of the same sample in three different states.”
The researchers demonstrated that the technique could decode the spatial arrangement of seven components within one structure. The results were published in Nature Communications.
“By using antibodies and other widely used agents to direct DNA probes to molecular targets, we might apply APR technology to decode the components and geometries of biological complexes,” says Schaus, who is currently a Wyss Institute Research Scientist. “The fact that individual DNA records carry unique, sequenceable barcodes and that the method is scalable may enable us to one day follow, individually, thousands, or millions of macromolecules in a biochemical pathway.”
As the work progressed, Schaus and his colleagues explored other applications based on the conditional DNA generation that they developed, termed ‘Primer Exchange Reactions’.
“I helped a graduate student in the lab, Josie Kishi, further characterize and develop the core APR reactions involving copying and releasing arbitrary DNA sequences, and she spearheaded a variety of applications,” Schaus says.
Delivering drugs, signaling the presence of disease in the body, and detecting toxins in the environment are all long-term potential applications for DNA-based molecular devices. And to create these devices, researchers often need to synthesize long, complex, and properly-folded DNA molecules to meet immediate needs.
The work completed in Peng Yin’s lab offers a new approach that automates the production of DNA segments and joins them in sequence according to a planned design. Called Primer Exchange Reaction (PER) cascades (see animation), the researchers used the method to engineer devices with diverse functions: manufacture of DNA-origami components, and DNA nanostructures that sense, amplify, record, or evaluate environmental signals. The work was published in Nature Chemistry.
“The core of DNA nanotechnology is the incredible programmability of interactions between DNA strands, based in part on enormous information content and base pairing, but also on the many ways technologists have devised for choreographing static and even dynamic interactions between them,” says Schaus. “When one adds one of the many DNA-acting enzymes that enable operations like cutting, ligation, and copying to the toolbox, the possibilities are enormous, exciting, and often surprising. This potential has led the Wyss Institute here at Harvard to launch the Molecular Robotics Initiative, providing funding and encouraging collaboration and translation. Apart from end goals resembling humanoid robots, the ‘robotics’ term serves as a motivating and exciting target to see how far we can push complex abilities at the molecular scale.”
Schaus and his colleagues plan to continue developing molecular tools. “Our goal from the beginning was to enable a single-molecule description of molecular interaction,” he says. “With unique barcodes applied to tens of thousands of individuals in a single experiment, and a next-gen sequencing readout, we hope to fulfill this need. We believe it will enable new views of cell function, much like single cell transcriptomics has. We also have twists on the APR mechanism that allow for other types of data to be recorded, and some exciting plans for PER applications as well.”
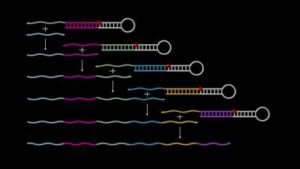
Primer Exchange Reaction (PER) cascades enable the autonomous growth of single-stranded DNAs. (Wyss Institute at Harvard University)